End of February I attended Analytics and Data Summit 2020 in Santa Clara where I presented 2 sessions and a hands-on lab. Delivering a hands-on lab is a double challenge: content and environment. Not only you have to prepare the content, including the lab with all the instructions, but you must also find a way to allow attendees to follow the lab in the simpler possible way (and avoiding the lab to crash and spend the whole time debugging).
In the old days it was generally a big virtual machine. Attendees had to copy from bunch of USB sticks being passed around, deploying it on their own laptop and crossing fingers it would work. That was the best way to lose half an hour, if not more, of the lab just doing the deployment and fixing issues.
Cloud allows to easily prepare lab environments
Nowadays, thanks to the various cloud offers, it’s easier to prepare environments which wouldn’t require attendees to install anything locally but only connect to existing remote instances.
This is also the solution I adopted to keep it as simple as possible for attendees: all they were required to have was a browser and an internet connection.
My hands-on lab was about creating and using a property graph in Oracle from a database source. A lab really starting from scratch and showing all the steps, which could be easily adopted even in a company environment to define and load graphs using the Oracle Database as storage (feel free to use the lab yourself if you are interested in the topic).
Translating my own sandbox into a more “enterprise” one
For all my own experiments I generally work on a sandbox I have at home on which I run Docker containers for everything: database, JupyterLab, Oracle Analytics Server etc.
Thinking at this lab in my own sandbox it is 2 containers: one for an Oracle Database (18c XE would be a perfect match) and another one for JupyterLab with PGX (Oracle Property Graph) installed and Python 3.
As you maybe saw from some of my old posts here, I generally work with graph from Python notebooks, connecting via JPype from Python to Java where the PGX client is executed. It isn’t the most natural approach for Oracle Property Graph, but the power of notebooks coupled with everything versioned in Git is priceless.
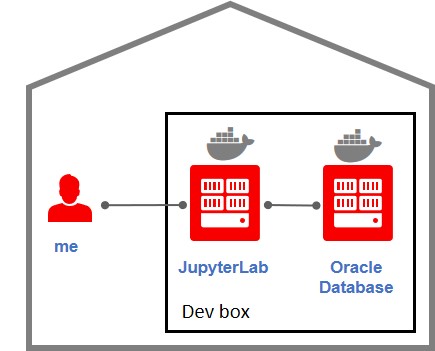
An extremely simple environment, no security, no firewalls, no SSL, no authentication: just the most simple and open sandbox. Using the IP address of my sandbox and the random port Docker assigned to the containers.
I don’t need security because the sandbox host is accessible only when connected to my home network (locally or via VPN).
If I would use that for the hands-on lab there is no chance to have attendees following and understanding things. Docker works on Windows and macOS, but not everybody has it. If people have a corporate laptop, they probably aren’t allowed to install anything for security reasons. If I wanted to use that I had to provide a virtual machine and inform attendees to install VirtualBox before to attend.
Attendees also would have had to learn Python, cx_Oracle for queries on the database via Python and JPype for the interaction with PGX: all things which aren’t the topic of the lab, but would easily take 75%, or more, of the time!
The challenge: keep it simple!
How to turn all that home-sandbox into something simpler? Ideally using only a browser?
This is where Oracle Cloud jumps in and allows you to deploy all that almost easily.
Autonomous Database
First step was replacing the Docker container for the database. That is the easier part: Oracle has various database services in its cloud offer.
For the hands-on lab I selected the Autonomous Data Warehouse service. Not only I count on the autonomous side of the service to not have to care or manage anything myself (it’s a lab anyway, there isn’t any tuning etc. to do), but it also comes with SQL Developer Web available by default.
The normal SQL Developer is just download an archive, extract, run. But if you can have the same in your browser it’s even better for a hands-on lab.
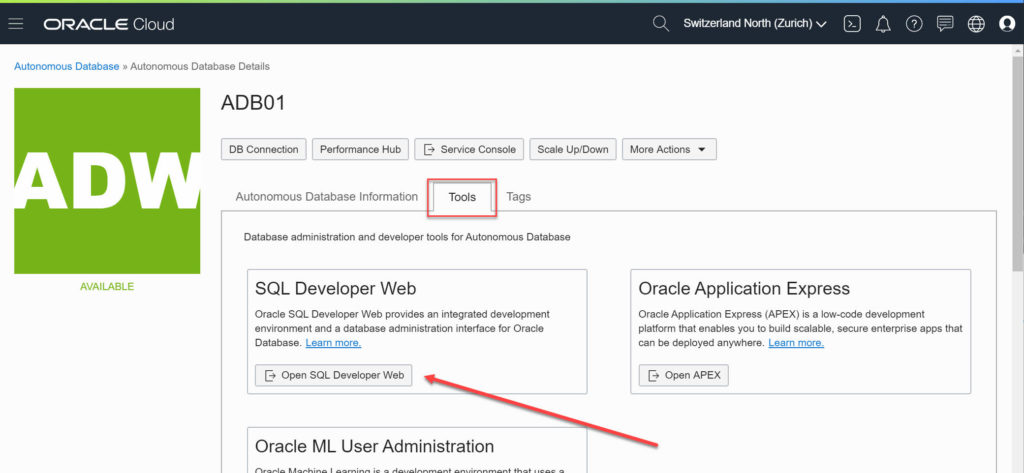
No need to download the Client Credentials (Wallet), to have the right Java etc. Just a browser is all it requires!
This solved my database part: every attendee would receive a unique username/password to connect and the URL for SQL Developer Web. To make it more user-friendly I used a proxy for it, so the URL is nicer (the one ADB gives you is really ugly!).
PGX JShell client made easy: JupyterLab with iJava kernel
To replace the Python notebooks using JPype I decided to go directly for the Oracle Graph thick client which can either be a Groovy based CLI or a JShell based CLI. In this way there is nothing else than property graph itself to learn, attendees will be directly on the product!
The annoying part is that a CLI isn’t the best environment for a lab: a black screen with limited run/edit/rerun options.
This is where Jupyter is really an awesome tool: somebody wrote a JShell kernel which can be deployed. This allows attendees to use the normal JShell CLI syntax and commands, but working in a notebook making things easier and more interactive with comments, cells of code which can be easily edited and re-executed etc.
The downside is that there is no JupyterLab service on Oracle Cloud! I would need an independent JupyterLab for every single attendee to the lab (separate sessions).
Oracle Data Science
To be precise there is one now. The Oracle Data Science service provides you with a JupyterLab environment. But it’s targeted toward machine learning, not really for being used as a user-friendly replacement for the Oracle Graph client.
It has Python 3 and various data manipulation, visualization and ML packages.While it is possible to extend it by installing other things, my lab need is really too far to justify using this service as a base.
Not a big deal, I already wrote on how to have JupyterLab setup in Oracle Cloud: https://gianniceresa.com/2019/12/jupyter-sandbox-in-an-oracle-cloud-compute-instance/
The only difference is that I need to provide that environment for up to 55 attendees, all using the Oracle Property Graph in memory engine at the same time. A single compute instance would provide a really poor user experience and be highly at risk of crashing for everybody in case of an issue.
This is why I decided to adopt smaller compute instances but many of them in parallel: 4 users per instance and 2 instances as spare in case of … (because Murphy’s law). This means a total of 16 compute instances.
Because having to deal with multiple users with independent sessions, I adopted JupyterHub, the multi-user JupyterLab environment. It will deal with authentication and provide every user with an isolated JupyterLab instance keeping things simple and clean.
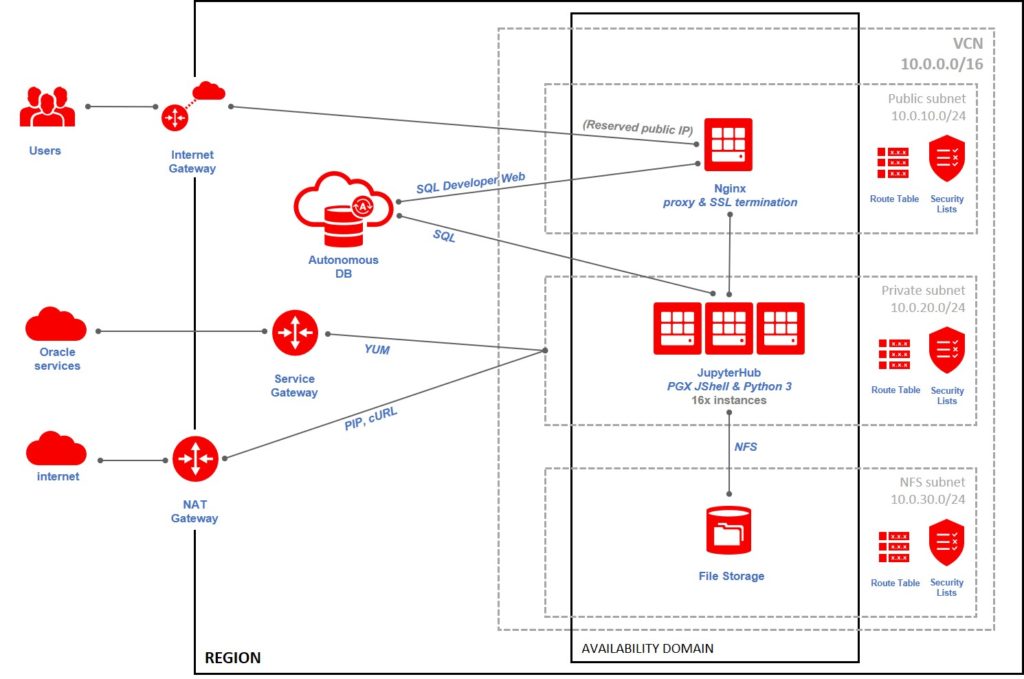
Security: a single public point of access to all the services over SSL
The final component of the setup is the “glue” for all the pieces. For security reasons I don’t want to expose all these instances and services publicly to the internet. I want a single point of access and everything over SSL connections as there are credentials sent around. This is why I adopted a proxy located in front of everything.
A single small compute instance running nginx acting as reverse proxy for SQL Developer Web, forwarding a friendly URL to the ugly one provided by Autonomous Database.
The same nginx acting as reverse proxy and SSL termination in front of all the JupyterHub instances (16 of them). I could have used load balancing in nginx to share the load across the 16 instances, or even the load balancing provided by OCI Compute, but I preferred to use 16 different URLs. In this way I could easily ask users of a faulty instance to move to another one by changing the URL they used.
nginx: what’s that?
You are probably familiar with, or know, the Apache HTTP Server. It has been the go-to web server for years and still is. It has various modules including one doing redirections and proxy.
nginx (pronounced engine x) is an HTTP and reverse proxy server, a mail proxy server, and a generic TCP/UDP proxy server, originally written by Igor Sysoev.
I first discovered it around 2005 when I was setting up some web server for a website with high traffic. Apache HTTP Server was struggling to scale. nginx did the job for a fraction of the memory and scaled in an unbelievable way. Since 2009-2010 it is more and more used, and nowadays it is the most used web server.
As compute instances can have changing IP addresses, I used a reserved public IP which I linked to various subdomain of gianniceresa.com. This also allowed me to safely provision the trusted SSL certificates (via Let’s Encrypt) upfront knowing the IP wouldn’t change.
To make sure the notebooks (files) of attendees wouldn’t be lost in case of change of instance, but also to share with all of them the lab documents and the wallet for the database connection, the 16 compute instances all mounted the same File Storage file system (an Oracle Cloud service providing a NFS shared folder).
The security was ensured by the firewall running on each compute instance, by the ACL on the Autonomous Database instance and by the security lists of the public and private subnets I created in my Virtual Cloud Network.
Oracle Cloud made it simple to setup and manage
I must admit it has been a really fun exercise. Designing the infrastructure, the networks, the security and finally implement it. Oracle OCI makes it really simple to build even the most enterprise-level complex setup with multiple networks, gateways, route tables etc. It just requires some design first to know what you want.
Ansible also played a role. No way I would install the 16 compute instances by hand, with the risk of messing up something because of a “human” copy/paste mistake. I setup Ansible on the compute instance acting as proxy, wrote my playbook and waited. With a single command, in about 1 hour, I had 16 perfectly identical compute instances fully setup with users, passwords, installed and configured tools, mounted shared folders. Everything!
All in all …
This isn’t a technical post, I just wanted to share how the cloud allows you to easily scale something you tested in private to a larger and stable setup which can be shared with multiple users.
This could easily translate to go from a single-user notebook solution for ML to a safe solution for a group of data scientist collaborating together.
In case you were wondering: the hands-on lab went very well! Not a single technical issue, nobody couldn’t do the lab because of not being able to have the required setup. Every unique credential given to users worked, the proxy did an awesome job. Both the Autonomous Database and the compute instances have been fully stable and not overloaded at any point in time. (At the same conference I assisted to a lab where they spent the whole lab just trying to connect because all the credentials expired. I’m quite happy all went well for me.)
